Heads Up!
You might not be aware, but Australia and New Zealand will be transitioning away from the Burrlioz software (and by extension – the default use of the Burr distribution) for SSD modelling. This has been motivated by two considerations:
- Burrlioz is now over 25 years old and many users are reporting that the software no longer runs on today’s modern computers
there have been new statistical developments (most importantly model averaging and mixture modelling) and software developments (R package ssdtools) for SSD modelling that are superior to Burrlioz.
and
This is an essential course if you are a practising ecotoxicologist who has an interest in:
- concentration-response (C-R) modelling
- species sensitivity distributions (SSDs)
- the Burrlioz software
- ssdtools software
- establishing guideline values
just learning about model averaging
or
Course description
The species sensitivity distribution (SSD) underpins the derivation of protective concentrations (PCx) that are applied in most current formal water quality guideline value (GV) derivations (Fox et al. 2021). The data underpinning the SSD are the toxicity estimates extracted from individual species concentration-response (CR) experimental data and should ideally represent no effect toxicity estimates.
A range of new statistical approaches is being developed and/or adopted in ecotoxicology, when combined, have the potential to greatly improve the estimation of no-effect toxicity values from Concentration-Response (CR) experimental data.
Commencing with with an introduction to CR modelling, the course will cover the most recent methods for estimating toxicity (and more specifically no-effect toxicity) from concentration response models using R.
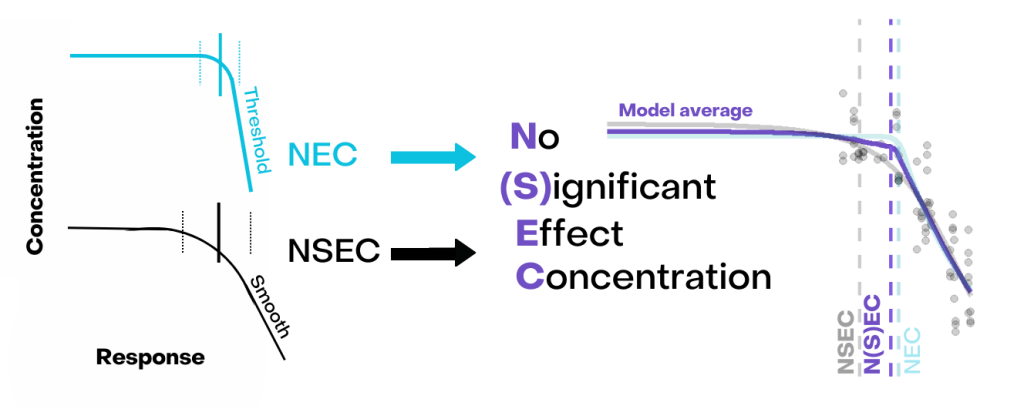
Topics to be covered include: the difference between linear, non-linear and threshold models; natural scaling of response variables (endpoints) and generalised modelling; and the distinction between derived toxicity metrics (NEC, ECx).
A new metric for estimating a no-effect toxicity, the no significant effect concentration (NSEC) will be introduced and methods for computing it using R explained. Ample examples and exercises will be used to demonstrate the application of CR models and the estimation of toxicity values in R, using frequentist approaches via the drc package, and within a Bayesian framework using the backage bayesnec. Finally we will discuss and demonstrate the advantages of using the model averaging framework for toxicity estimating using CR data.
Learning outcomes
On completion of this course you will:
Recognition
If you choose, you may take an online assessment on completion of this course to test your understanding of C-R modelling. The assessment will be a combination of multiple-choice questions together with demonstration of competency using R and various packages and tools to perform various calculations associated with fitting and using C-R models. The assessment is not difficult if you have worked through the course and satisfactorily completed the quizzes along the way. A pass of at least 70% is required to qualify for an ECOTOX-U certificate of completion in C-R modelling.
Background reading
The modelling covered in this course will be based extensively on the use of drc and bayesnec.
- The drc helpfile contains considerable information it’s use, with the associated publication covering many of the important details here.
- A simple tutorial for dose-response modelling in R using drc can be found here.
- A simple tutorial for dose-response modelling in R using drc can be found here.
- For bayesnec, there is substantial background information and examples on the github website, including:
- single model usage, multi-model usage,
- detailed information on the available models,
- information on priors, and
- using posteriors for comparing across model sets.
- NSEC Information on the new metric for estimating a no-toxicity, the no significant effect concentration (NSEC) is detailed in this recent publication.
- A newly developed tool for estimating NSEC values from bayesnec and drc models can be found here.
Software installation
The course will use the R packages drc and bayesnec, run through RStudio. Please make sure you have these packages/programs running on your individual laptop prior to the in-person event in Townsville.
Course notes - will be provided on the day.